Frequently Asked Questions
What are the current barriers to others accessing data within the Cambridge Biomedical Campus?
Cambridge-based researchers are already able to ask other researchers for data, based on their experience and knowledge of what is available. This information is then sent in a downloaded format to approved researchers, requiring a lot of computer space and power. CYNAPSE will make it much simpler for approved researchers to access research data safely and securely, without the need for sharing or downloading original datasets. The CYNAPSE Service Delivery Team will ensure any proposed access is safe and appropriate.
Examples of datasets on the platform can be found on the Studies page.
Access and Data
Can research teams based outside of the Cambridge Biomedical Campus contribute to the database?
CYNAPSE initially focused on researchers based on the Cambridge Biomedical Campus. Several different specialities and departments participated in Phase 1 of the programme (building and testing the new platform). CYNAPSE is now open to additional groups across the University of Cambridge.
Can we invite study collaborators from outside of the Cambridge Biomedical Campus to use our workspace within CYNAPSE?
Academics from other institutions can be granted access to a researcher workspace, if the appropriate IG approvals are in place, after discussion with the Workspace Principal Investigator. Collaborations with industry are not yet available, pending the development of an appropriate costing model. To learn more about the different workspaces that will be available in CYNAPSE, visit the Patients & Public Governance page.
Will researchers be able to access 'live' data (data as it is being introduced into the system) or archived data (data that has been stored after permissions have been passed)?
As soon as 'sharable' data is uploaded into CYNAPSE, researchers will be able to apply for access to it. Once they have received the relevant approvals, they will be granted access. Only data that has valid permissions will be stored within and uploaded into CYNAPSE. If any relevant permission expires, the data will be disposed of in accordance with the original requirements of that particular study.
Are there plans to add central datasets like UK Biobank to the platform for all groups to use?
This is not yet available but is in the long term plan for CYNAPSE. IG agreements will need to be put in place with the data providers and a costing model agreed to ensure the large storage costs for these resources will be covered.
Will federation be available on the platform?
Part of the CYNAPSE programme will allow for connectivity between datasets amongst different research organisations without the need to move datasets to new locations (known as 'federation'). We will do this by developing technology to allow datasets held in these independent locations to be analysed simultaneously. The results of the separate analyses can then be combined without the original data ever having to move.
We began this process as part of the 'Multi-party trusted research environment federation: Establishing infrastructure for secure analysis across different clinical-genomic datasets' project. THis was one of nine short-term projects funded by UK Research & Innovation as part of Phase 1 of the DARE UK (Data and Analytics Research Environments UK) programme, delivered in partnership with Health Data Research UK (HDR UK) and ADR UK (Administrative Data Research UK). This proof of concept project tested whether researchers could access and simultaneously analyse data that is held in the two protected areas of CYNAPSE and Genomics England. We plan to expand this to other areas in the future.
What information does the current data contain, and where does it come from?
The data is generated from participants who took part in research studies and contains:
- 'omic data (for example, DNA, RNA, or protein analysis)
- de-identified phenotypic datasets from previous or current research projects (such as age band, sex, ethnicity)
Some of the data in CYNAPSE were generated through pre-clinical research (this means using human cell lines or animal cells rather than research with human participants).
Can we add all types of data to the platform, including clinical data?
The platform has been predominantly created for genomic data but we are working with Lifebit to determine the platform and security enhancements required for the platform to work with all data types.
Does CYNAPSE contain data from patient records?
No. The human data in CYNAPSE comes from large research studies from consenting participants or from pre-clinical research. However, there are plans to add access to data from patient records through CYNAPSE in future. Where this happens, all information from patients will be combined (known as 'aggregated') with all information that could identify an individual removed (known as 'de-identified') to protect privacy.
Will this add to the data available for systematic review studies?
Not in terms of the actual source data. Any resulting publications & results could be included.
What are the consequences of misuses of the data?
Researchers accessing data, and often their organisations, sign an agreement that sets out the conditions of use. Breaking these terms could result in legal, financial, reputational, and likely employment consequences.
A copy of the Acceptable Use Policy can be found here.
Will anyone be able to contribute data to CYNAPSE?
Researchers or clinicians approved by the University of Cambridge Clinical School and the CYNAPSE IG team will be able to contribute research data to CYNAPSE.
Who is responsible for ensuring only relevant users have access to a dataset?
Only the CYNAPSE Service Delivery team, which includes an Information Governance advisor, will have permissions to grant access to a workspace (and the datasets within it), making sure that researchers have provided details of the proposed research project, alongside NHS Research Ethics (where necessary) and Research & Development approval.
See also Governance.
What happens if a researcher leaves a research group, will they still be able to access data?
If the researcher is no longer employed by the University of Cambridge, their access will be revoked. It should be noted that in some cases researchers will move to a visiting worker/collaboration state and retain access.
Are you confident the data will be secure from hackers?
We use multiple layers of protection to secure the data from hackers, including:
- All infrastructure resources are placed in a secure private network and access to this network is only accessible through a virtual private network (VPN)
- All infrastructure services, including storage have secure protocols and encryption (can only be read by those with the correct key)
- When data is 'at rest' (not currently being accessed or used) encryption and access control is enforced
- Annual 'penetration testing' by an independent provider, which involves a simulation of real-world attacks by authorised security professionals in order to find potential weaknesses in security
- All storage on the platform has equivalent encryption, AES-256, both at rest and in in-flight
- Ingest data buckets are accessible outside of the platform via credentials provided to members of the Service Delivery team
- All data buckets generated by the platform (project, analysis results) are only accessible within the environment
A simple version of our high-level architecture can be found below:
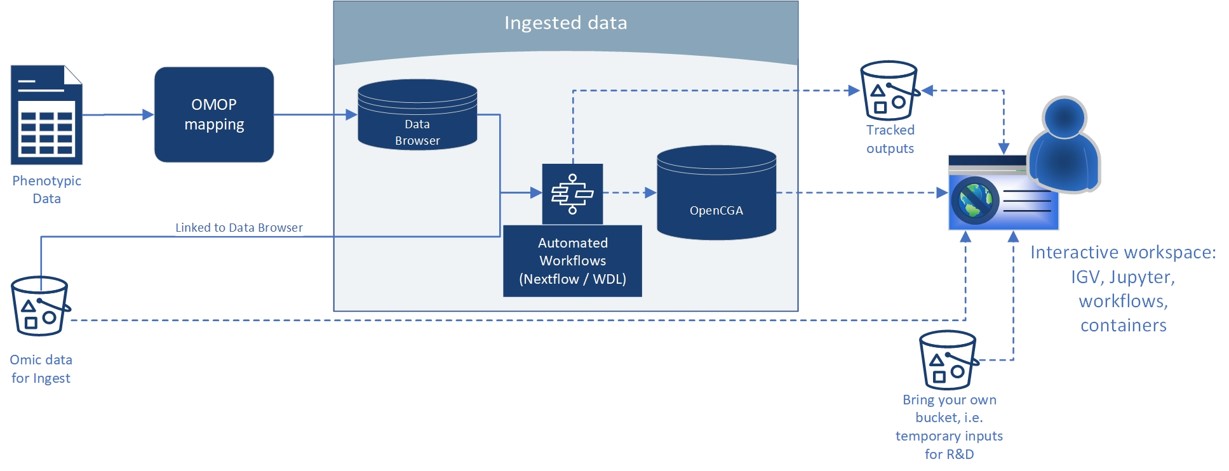
*Intelligent storage: If data is not used for a period of time it automatically migrates to lower performance and lower cost storage. Frequent access will result in the data migrating to higher speed tiers, and costs increasing. Over 90% of data generated in genomics is infrequently accessed.
Governance
Who will be the owner of CYNAPSE? Who else is involved in the project?
The University of Cambridge is the owner of the system. Health Innovation East (previously known as Eastern AHSN) and Cambridge University Health Partners were been jointly commissioned to project manage the implementation of the software platform and cloud-based TRE (Lifebit CloudOS) provided by our technology partner Lifebit.
Who decides what data can be stored in CYNAPSE? Who is responsible for ensuring data has the correct consent?
The data are generated by individual researchers, who are legally responsible for the safe storage and use of the datasets. Any data for inclusion in CYNAPSE will be reviewed to ensure that it meets a minimum set of standards to ensure that it is in a format that is useful to others before it is uploaded. The CYNAPSE Information Governance team are in place to develop this process alongside the CYNAPSE Co-creation Group and will ensure all Information Governance and data access procedures are followed, check where the data has come from, ensure it has the right approvals and the correct consent in place (or has no need for consent). See also Patients & Public Governance page and Data Users Governance page.
Will patients be involved in the governance of the programme?
Yes, as well as our previous work with the CYNAPSE Co-creation Group, we have a patient representative on the CYNAPSE Steering Committee, Programme Board, and on a number of workstreams. See also Patients & Public landing page.
Other
What does CYNAPSE stand for?
CYber ceNtre for secure data Analytics and Patient-directed reSEarch
What are TREs?
TREs are secure computing environments that hold data, allowing approved researchers to access and analyse information to support scientific studies. You can read more about TREs here.
Will the partnership with Lifebit last for the whole CYNAPSE programme?
A 4-year contract has been signed with Lifebit to provide the environment after which time the programme will be reviewed before any further contracts are signed.
What will the ongoing costs be for my team to use the platform?
Forecasted costs for storage and compute can be provided. Please get in touch via the help desk or at cynapse@healthinnovationeast.co.uk to discuss.
What tools are available on the platform and can we add our own?
For interactive analysis the platform provides Jupyter notebooks with language support for R, Python and Bash. Researchers also have access to the terminal, can install packages, pull code from repositories and use Docker images. Many common bioinformatic tools are available out of the box, but instance images can be generated with specific requirements.
For pipeline analysis, you are able to use Nextflow or Cromwell/WDL pipelines provided these declare Docker images. Private repository access is also possible.
What is FedPaaS and how is it relevant to CYNAPSE?
FedPaaS stands for Federated Platform as a Service. This term means that cloud infrastructure that is common across multiple users is shared. Moving to a FedPaaS system means that costs can be reduced through sharing the cost of common functionality. In the context of CYNAPSE, functionality like an airlock and compute methods are shared through Lifebit's FedPaaS platform. CYNAPSE data is still stored in CYNAPSE's AWS accounts and protected by the CYNAPSE firewall. Therefore, there is no change in the security of the data.